Applied graph analytics for indication expansion in drug discovery.
Part 2 - Graph Patterns for Indication Selection
This is a continuation of a multi-part series on Graph Patterns for Indication Selection.
Part 1 - Patient Recruitment using Graphs
Introduction
Indication expansion aims to increase the value of your drug by finding additional diseases that can benefit from it. Typically the drug is already in clinic or in market. More indications = more money. This analysis tells you “what” diseases makes sense based on biological principles. Another type of analysis known as indication sequencing describes the “how” and “when” to expand strategically.
All therapeutics, at a very fundamental level, basically translates to “give a substance to a biological thing to modulate a biological event that, when it is abnormal, is part of some disease”.
The key intuition here is to think of the association between a drug and the indication as a function of the biological effects it can modulate within a biological context. For example, you take an Advil when you have a migraine but Advil treats inflammation, which happens to be part of the symptoms of a migraine.
What exactly is a biological event?
A biological event refers to any measurable change within a living system caused by a specific interaction, such as the activation or inhibition of a biochemical pathway, receptor, enzyme, or gene.
How the substance interacts with the target to exert the correct change to a biological event is what we refer to as the mechanism of action (MOA). Biological complexity gives us an opportunity to exploit fundamental processes in different ways through a variety of targets including genes, proteins, protein-protein interactions, specific nucleotides, epigenetic modifications, and so on.
The participants in these effects vary widely, including proteins, small molecules, macro-complexes, and entire tissues. Biological effects also depend on context: the same molecule may behave differently in distinct tissues or under varying conditions, such as stress or infection.
The use of graphs in indication expansion is well-known. To understand why graphs are so powerful for this type of work, we will trace Dupilumab, a biologic that was initially approved for atopic dermatitis (AD) but has since gained approvals across multiple chronic inflammatory diseases such as COPD and asthma.
Graphing Biology
Dupilumab is a monoclonal antibody that acts as a receptor antagonist by binding to the alpha-subunit of interleukin-4 receptor (IL4R).
I’ve actually seen knowledge graphs in the wild that look like this.
This is useless. It gives you no analytical advantage, regardless of how “expertly curated” it is. The graph topology does not enable you to make deductions or inferences. Most of the value in your graph is in the design of the ontology.
Here’s what you should do instead.
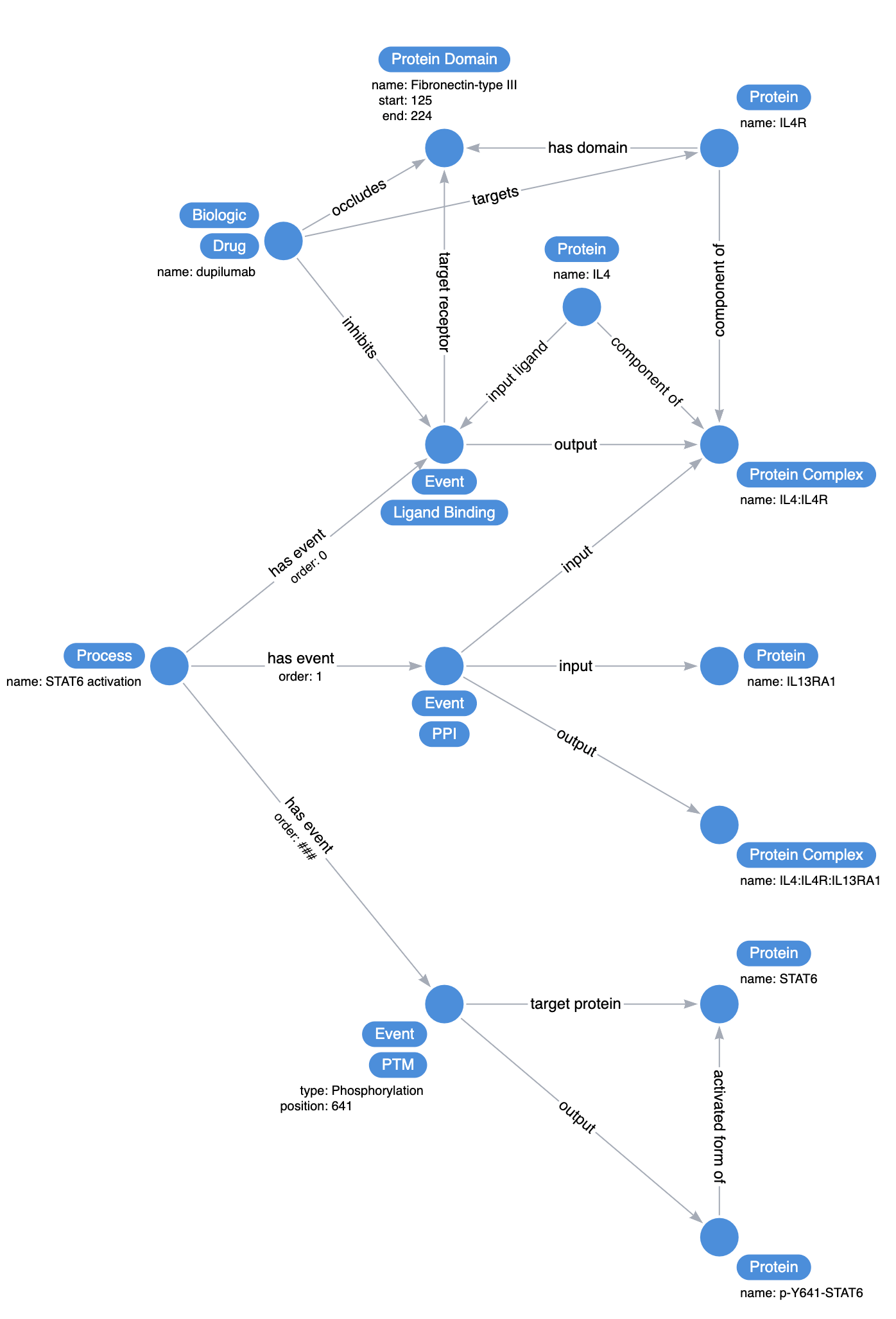
Use a concept called Event that captures useful information about biological reactions such as what the input substrate, output and the context of the event.
Next, describe the specific event that the drug is affecting. Dupilumab binds to IL4R and occludes the fibronectin domain, preventing IL4 from binding.
Next, we introduce the Process concept to assign the logical order to a collection of events. Zooming out and expanding the diagram above:
Dupilumab works because it attenuates Th2 pathway activation by crippling STAT6 activation. We can map the impact of the inhibition through evaluating the transcriptional targets of STAT6. To find these targets, you can rely on literature mining or determine these target empirically using epigenetic data sources such as ENCODE. Among these targets include CCL13, IGHE, IGHG1, and IGHG4 that drive type 2 inflammation. Importantly, STAT6 activation leads to IL4R upregulation and increase IL4 secretion, and executes a positive feedback loop to further amplify the response.
Causal Biology Modelling & Inference
Dupilumab is a blockbuster drug with 6 approved indications and more in the pipeline. This “pipeline in a product” type drug is extremely profitable because the drug targets a specific mechanism that is shared across a collection of diseases, in this case, STAT6 activation in type II immune response related diseases.
What happens when STAT6 is activated? What phenotypes are expected? What diseases are relevant?
We can leverage graph algorithms to answer these questions.
First, let’s build and visualize the network.
Starting from STAT6, mark all its transcriptional targets using ChIP-seq evidence. Specifically, you are looking for co-localization of ChIP-seq peaks for STAT6, H3K4me3, and H3K27Ac.
For any STAT6 targets, that are known to have DNA binding domains or are known transcription factors, expand out the transcriptional targets.
Repeat the process 2-3 times outwards
For all the genes marked - load in disease associations
Apply circular drawing algorithm for visualization
The positions of nodes in this projection are not arbitrary - they are determined by the topology of the network. Nodes that are more closely related (for example, those that share many edges or are part of the same community) tend to be placed next to one another. You are essentially seeing a one‐dimensional (angular) projection of the network’s connectivity. Nodes placed next to each other are likely to interact or be functionally related, while nodes that are far apart on the circle are less directly connected
In this application, we are doing a sanity check to see if the projection aligns with what we’d expect to see. For example, we know that Dupilumab is indicated for atopic eczema, so we should expect to find it somewhere here in this network. Indeed, we find it placed alongside CLDN1 and a variety of skin related disorders.
So far so good.
An interesting cluster with high convergence around STAT3 and a collection of chronic inflammatory diseases such as ulcerative colitis, psoriasis, and IBD.
The connection become clear when we run a path-finding algorithm between STAT3 and STAT6. Turns out, STAT6 drives the expression of IL4R and IL4 secretion. IL4R in heterodimer with IL13RA1 or IL2RG are both capable of phosphorylating STAT3, representing an alternate mechanism outside of the canonical IL-6 family activation loops. I’d suspect that Regeneron will starting going after treatment refractory UC and IBD soon.
Assessing efficacy for strategic expansion
“Pipeline in a product” are the holy-grail products that every pharma is chasing. When staging the launch sequence, one of the successful ways to build momentum is to use a narrow-first approach. Among the different variables for optimizing this launch strategy, one of them is to ensure you line up indications that you know are going to demonstrate strong efficacy and high market differentiation from standard of care. Translated into systems biology, we can interpret this as saying, does our drug impact the causal biological process the most.
How do you quantify that? Walk through the graph and calculate the path weights. (more on this in part 3 of this series)
To illustrate this, let’s use the recent example of BMS’s Cendakimab dropping out against Dupilumab. This was a wise decision because Cendakimab would have 100% lost this fight.
Cendakimab is a IL13 inhibitor. The goal is still to decrease IL4/IL13 signaling, which is the same as saying STAT6 suppression. There critical event is STAT6 phosphorylation, which happens under IL4R-alpha dependent heterodimerization. We can kick this cascade off at these entry events:
IL4 binds to IL4R → recruits IL13RA1
IL13 binds to IL4R → recruits IL13RA1
IL13 binds to IL13RA1 → recruits IL4R
At first glance, it might look like an IL13 blockade alone could achieve meaningful, or at least, comparable results because it serves as 2 potential entry points. This argument falls apart when we include the biological context in which these events are taking place. Specifically, IL13 binds to IL13RA2 (a decoy receptor) with much higher affinity than IL13RA1. Also, IL4 is still able to exert its effect, independent of IL13 blockade.
At best, Cendakimab might edge out a win on dosing regiment efficiency and a narrower ADR profile, but it’s not enough to drive differentiation and improvement over the current standard of care.
Conclusion
In this post, we dived deeper into how network graph analytics can help inform critical decisions made along the drug discovery landscape. Building a model to capture the complexity of systems biology can only be done in a structured ontology and knowledge graph. The investment in this resource unlocks powerful capabilities that research teams can use to answer questions in indication selection and strategic expansion in a principled way.
What’s Next?
Upcoming in this series, we’ll switch gears into building scoring models and graph neural networks that can help take safety risks off the table when evaluating indications.
BioBox is the knowledge infrastructure for modern biopharma research, built for drug hunters who need to integrate multi-modal data, engineer knowledge, and test hypotheses at scale. To learn more, please visit our website at https://biobox.io or click here to contact us.